1 c1i32D_
100.0
51
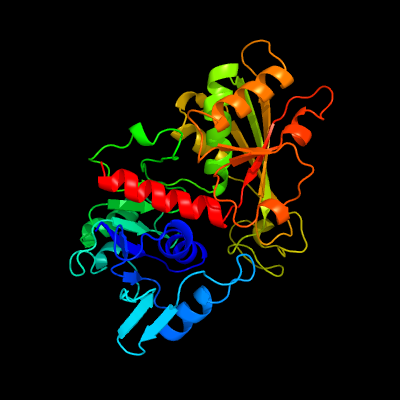
PDB header: oxidoreductaseChain: D: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase;PDBTitle: leishmania mexicana glyceraldehyde-3-phosphate2 dehydrogenase in complex with inhibitors
2 c5ld5C_
100.0
53
PDB header: oxidoreductaseChain: C: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystal structure of a bacterial dehydrogenase at 2.19 angstroms2 resolution
3 c2ep7B_
100.0
60
PDB header: oxidoreductaseChain: B: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: structural study of project id aq_1065 from aquifex aeolicus vf5
4 c3docD_
100.0
57
PDB header: oxidoreductaseChain: D: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase;PDBTitle: crystal structure of trka glyceraldehyde-3-phosphate dehydrogenase2 from brucella melitensis
5 c1obfO_
100.0
51
PDB header: glycolytic pathwayChain: O: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase;PDBTitle: the crystal structure of glyceraldehyde 3-phosphate2 dehydrogenase from alcaligenes xylosoxidans at 1.7 a3 resolution.
6 c3hq4R_
100.0
52
PDB header: oxidoreductaseChain: R: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase 1;PDBTitle: crystal structure of c151s mutant of glyceraldehyde-3-phosphate2 dehydrogenase 1 (gapdh1) complexed with nad from staphylococcus3 aureus mrsa252 at 2.2 angstrom resolution
7 c1rm4O_
100.0
55
PDB header: oxidoreductaseChain: O: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase a;PDBTitle: crystal structure of recombinant photosynthetic glyceraldehyde-3-2 phosphate dehydrogenase a4 isoform, complexed with nadp
8 c6ok4A_
100.0
51
PDB header: oxidoreductaseChain: A: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystal structure of glyceraldehyde-3-phosphate dehydrogenase (gapdh)2 from chlamydia trachomatis with bound nad
9 c2pkrI_
100.0
55
PDB header: oxidoreductaseChain: I: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase aor;PDBTitle: crystal structure of (a+cte)4 chimeric form of photosyntetic2 glyceraldehyde-3-phosphate dehydrogenase, complexed with nadp
10 c3b20R_
100.0
54
PDB header: oxidoreductaseChain: R: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase (nadp+);PDBTitle: crystal structure of glyceraldehyde-3-phosphate dehydrogenase2 complexed with nadfrom synechococcus elongatus"
11 c1cerC_
100.0
61
PDB header: oxidoreductase (aldehyde(d)-nad(a))Chain: C: PDB Molecule: holo-d-glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: determinants of enzyme thermostability observed in the2 molecular structure of thermus aquaticus d-glyceraldehyde-3 3-phosphate dehydrogenase at 2.5 angstroms resolution
12 c2d2iO_
100.0
54
PDB header: oxidoreductaseChain: O: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase;PDBTitle: crystal structure of nadp-dependent glyceraldehyde-3-2 phosphate dehydrogenase from synechococcus sp. complexed3 with nadp+
13 c1hdgO_
100.0
58
PDB header: oxidoreductase (aldehy(d)-nad(a))Chain: O: PDB Molecule: holo-d-glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: the crystal structure of holo-glyceraldehyde-3-phosphate dehydrogenase2 from the hyperthermophilic bacterium thermotoga maritima at 2.53 angstroms resolution
14 c3hjaB_
100.0
58
PDB header: oxidoreductaseChain: B: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystal structure of glyceraldehyde-3-phosphate dehydrogenase from2 borrelia burgdorferi
15 c2b4rQ_
100.0
53
PDB header: oxidoreductaseChain: Q: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystal structure of glyceraldehyde-3-phosphate dehydrogenase from2 plasmodium falciparum at 2.25 angstrom resolution reveals intriguing3 extra electron density in the active site
16 c1ihxD_
100.0
51
PDB header: oxidoreductaseChain: D: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase;PDBTitle: crystal structure of two d-glyceraldehyde-3-phosphate2 dehydrogenase complexes: a case of asymmetry
17 c3cieC_
100.0
51
PDB header: oxidoreductaseChain: C: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystal structure of glyceraldehyde 3-phosphate2 dehydrogenase from cryptosporidium parvum
18 c3h9eO_
100.0
50
PDB header: oxidoreductaseChain: O: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase, testis-specific;PDBTitle: crystal structure of human sperm-specific glyceraldehyde-3-phosphate2 dehydrogenase (gapds) complex with nad and phosphate
19 c2x5kO_
100.0
43
PDB header: oxidoreductaseChain: O: PDB Molecule: d-erythrose-4-phosphate dehydrogenase;PDBTitle: structure of an active site mutant of the d-erythrose-4-phosphate2 dehydrogenase from e. coli
20 c5j9gB_
100.0
48
PDB header: oxidoreductaseChain: B: PDB Molecule: glyceraldehyde-3-p dehydrogenase;PDBTitle: structure of lactobacillus acidophilus glyceraldehyde-3-phosphate2 dehydrogenase at 2.21 angstrom resolution
21 c4qx6A_
not modelled
100.0
54
PDB header: oxidoreductaseChain: A: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase;PDBTitle: crystal structure of glyceraldehyde-3-phosphate dehydrogenase from2 streptococcus agalactiae nem316 at 2.46 angstrom resolution
22 c5ur0B_
not modelled
100.0
50
PDB header: oxidoreductaseChain: B: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystallographic structure of glyceraldehyde-3-phosphate dehydrogenase2 from naegleria gruberi
23 c1s7cA_
not modelled
100.0
52
PDB header: structural genomics, oxidoreductaseChain: A: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase a;PDBTitle: crystal structure of mes buffer bound form of glyceraldehyde 3-2 phosphate dehydrogenase from escherichia coli
24 c4dibF_
not modelled
100.0
55
PDB header: oxidoreductaseChain: F: PDB Molecule: glyceraldehyde 3-phosphate dehydrogenase;PDBTitle: the crystal structure of glyceraldehyde-3-phosphate dehydrogenase from2 bacillus anthracis str. sterne
25 c3sthA_
not modelled
100.0
51
PDB header: oxidoreductaseChain: A: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystal structure of glyceraldehyde-3-phosphate dehydrogenase from2 toxoplasma gondii
26 c2i5pO_
not modelled
100.0
50
PDB header: oxidoreductaseChain: O: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase 1;PDBTitle: crystal structure of glyceraldehyde-3-phosphate2 dehydrogenase isoform 1 from k. marxianus
27 c2gd1P_
not modelled
100.0
61
PDB header: oxidoreductase(aldehyde(d)-nad(a))Chain: P: PDB Molecule: apo-d-glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: coenzyme-induced conformational changes in glyceraldehyde-3-2 phosphate dehydrogenase from bacillus stearothermophillus
28 c5jyfB_
not modelled
100.0
57
PDB header: oxidoreductaseChain: B: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: structures of streptococcus agalactiae gbs gapdh in different2 enzymatic states
29 c2yyyB_
not modelled
100.0
18
PDB header: oxidoreductaseChain: B: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystal structure of glyceraldehyde-3-phosphate2 dehydrogenase
30 c1cf2Q_
not modelled
100.0
21
PDB header: oxidoreductaseChain: Q: PDB Molecule: protein (glyceraldehyde-3-phosphate dehydrogenase);PDBTitle: three-dimensional structure of d-glyceraldehyde-3-phosphate2 dehydrogenase from the hyperthermophilic archaeon methanothermus3 fervidus
31 d1rm4a2
not modelled
100.0
62
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
32 c1b7gO_
not modelled
100.0
17
PDB header: oxidoreductaseChain: O: PDB Molecule: protein (glyceraldehyde 3-phosphate dehydrogenase);PDBTitle: glyceraldehyde 3-phosphate dehydrogenase
33 d3cmco2
not modelled
100.0
62
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
34 d1gado2
not modelled
100.0
56
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
35 d2b4ro2
not modelled
100.0
57
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
36 d1u8fo2
not modelled
100.0
56
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
37 d2g82a2
not modelled
100.0
62
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
38 d2pkqo2
not modelled
100.0
62
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
39 d1ggaa2
not modelled
100.0
53
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
40 d3gpdg2
not modelled
100.0
56
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
41 d1i32a2
not modelled
100.0
53
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
42 d1dssg2
not modelled
100.0
57
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
43 d1k3ta2
not modelled
100.0
52
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
44 d1obfo2
not modelled
100.0
54
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
45 c2czcD_
not modelled
100.0
17
PDB header: oxidoreductaseChain: D: PDB Molecule: glyceraldehyde-3-phosphate dehydrogenase;PDBTitle: crystal structure of glyceraldehyde-3-phosphate dehydrogenase from2 pyrococcus horikoshii ot3
46 d1hdgo2
not modelled
100.0
62
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
47 d1ggaa1
not modelled
100.0
43
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
48 d1k3ta1
not modelled
100.0
41
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
49 d3cmco1
not modelled
100.0
56
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
50 d1rm4a1
not modelled
100.0
44
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
51 d1hdgo1
not modelled
100.0
49
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
52 d1gado1
not modelled
100.0
45
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
53 d1i32a1
not modelled
100.0
39
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
54 d2g82a1
not modelled
100.0
56
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
55 d1u8fo1
not modelled
100.0
41
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
56 d1vc2a1
not modelled
100.0
54
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
57 d3gpdg1
not modelled
100.0
39
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
58 d1j0xo1
not modelled
100.0
41
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
59 d1obfo1
not modelled
100.0
46
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
60 d2pkqo1
not modelled
100.0
43
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
61 d2b4ro1
not modelled
100.0
47
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
62 d1dssg1
not modelled
100.0
39
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
63 c2qz9B_
not modelled
100.0
19
PDB header: oxidoreductaseChain: B: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: crystal structure of aspartate semialdehyde dehydrogenase2 ii from vibrio cholerae
64 c4dpkB_
not modelled
100.0
17
PDB header: oxidoreductaseChain: B: PDB Molecule: malonyl-coa/succinyl-coa reductase;PDBTitle: structure of malonyl-coenzyme a reductase from crenarchaeota
65 c2ep5B_
not modelled
100.0
18
PDB header: oxidoreductaseChain: B: PDB Molecule: 350aa long hypothetical aspartate-semialdehydePDBTitle: structural study of project id st1242 from sulfolobus tokodaii strain7
66 c2yv3B_
not modelled
100.0
23
PDB header: oxidoreductaseChain: B: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: crystal structure of aspartate semialdehyde dehydrogenase from thermus2 thermophilus hb8
67 c2g17A_
not modelled
100.0
17
PDB header: oxidoreductaseChain: A: PDB Molecule: n-acetyl-gamma-glutamyl-phosphate reductase;PDBTitle: the structure of n-acetyl-gamma-glutamyl-phosphate reductase from2 salmonella typhimurium.
68 c1ys4A_
not modelled
100.0
20
PDB header: oxidoreductaseChain: A: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: structure of aspartate-semialdehyde dehydrogenase from methanococcus2 jannaschii
69 c3uw3A_
not modelled
100.0
17
PDB header: oxidoreductaseChain: A: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: crystal structure of an aspartate-semialdehyde dehydrogenase from2 burkholderia thailandensis
70 c2gz3D_
not modelled
100.0
17
PDB header: oxidoreductaseChain: D: PDB Molecule: aspartate beta-semialdehyde dehydrogenase;PDBTitle: structure of aspartate semialdehyde dehydrogenase (asadh) from2 streptococcus pneumoniae complexed with nadp and aspartate-3 semialdehyde
71 c2q49B_
not modelled
100.0
15
PDB header: oxidoreductaseChain: B: PDB Molecule: probable n-acetyl-gamma-glutamyl-phosphate reductase;PDBTitle: ensemble refinement of the protein crystal structure of gene product2 from arabidopsis thaliana at2g19940
72 c3kubA_
not modelled
100.0
20
PDB header: oxidoreductaseChain: A: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: crystal structure of aspartate semi-aldehyde dehydrogenase complexed2 with glycerol and phosphate of mycobacterium tuberculosis h37rv
73 c1mb4B_
not modelled
100.0
18
PDB header: oxidoreductaseChain: B: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: crystal structure of aspartate semialdehyde dehydrogenase from vibrio2 cholerae with nadp and s-methyl-l-cysteine sulfoxide
74 c1t4bB_
not modelled
100.0
18
PDB header: oxidoreductaseChain: B: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: 1.6 angstrom structure of esherichia coli aspartate-2 semialdehyde dehydrogenase.
75 d1cf2o1
not modelled
100.0
20
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
76 c5jw6A_
not modelled
100.0
16
PDB header: oxidoreductaseChain: A: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: cystal structure of aspartate semialdehyde dehydrogenase from2 aspergillus fumigatus
77 c5cefA_
not modelled
100.0
17
PDB header: oxidoreductaseChain: A: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: cystal structure of aspartate semialdehyde dehydrogenase from2 cryptococcus neoformans
78 c4wojB_
not modelled
100.0
17
PDB header: oxidoreductaseChain: B: PDB Molecule: aspartate semialdehyde dehydrogenase;PDBTitle: aspartate semialdehyde dehydrogenase from francisella tularensis
79 c2ozpA_
not modelled
100.0
15
PDB header: oxidoreductaseChain: A: PDB Molecule: n-acetyl-gamma-glutamyl-phosphate reductase;PDBTitle: crystal structure of n-acetyl-gamma-glutamyl-phosphate reductase2 (ttha1904) from thermus thermophilus
80 c4zhsC_
not modelled
100.0
16
PDB header: oxidoreductaseChain: C: PDB Molecule: aspartate semialdehyde dehydrogenase;PDBTitle: crystal structure of aspartate semialdehyde dehydrogenase from2 trichophyton rubrum
81 c2hjsA_
not modelled
100.0
16
PDB header: structural genomics, unknown functionChain: A: PDB Molecule: usg-1 protein homolog;PDBTitle: the structure of a probable aspartate-semialdehyde dehydrogenase from2 pseudomonas aeruginosa
82 c1vknC_
not modelled
100.0
14
PDB header: oxidoreductaseChain: C: PDB Molecule: n-acetyl-gamma-glutamyl-phosphate reductase;PDBTitle: crystal structure of n-acetyl-gamma-glutamyl-phosphate reductase2 (tm1782) from thermotoga maritima at 1.80 a resolution
83 c3hskB_
not modelled
100.0
20
PDB header: oxidoreductaseChain: B: PDB Molecule: aspartate-semialdehyde dehydrogenase;PDBTitle: crystal structure of aspartate semialdehyde dehydrogenase with nadp2 from candida albicans
84 c2i3aD_
not modelled
100.0
12
PDB header: oxidoreductaseChain: D: PDB Molecule: n-acetyl-gamma-glutamyl-phosphate reductase;PDBTitle: crystal structure of n-acetyl-gamma-glutamyl-phosphate reductase2 (rv1652) from mycobacterium tuberculosis
85 d2gz1a1
not modelled
99.9
20
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
86 d1pqua2
not modelled
99.9
17
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
87 d1t4ba2
not modelled
99.9
16
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
88 d1mb4a2
not modelled
99.9
19
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
89 d2gz1a2
not modelled
99.9
19
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
90 d2czca2
not modelled
99.8
17
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
91 d1cf2o2
not modelled
99.7
24
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
92 d1b7go2
not modelled
99.7
19
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
93 d2hjsa2
not modelled
99.7
17
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
94 d2hjsa1
not modelled
99.7
12
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
95 d2cvoa1
not modelled
99.6
18
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
96 d2g17a1
not modelled
99.6
18
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
97 d1vkna1
not modelled
99.6
15
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
98 d2czca1
not modelled
99.6
20
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
99 d1b7go1
not modelled
99.5
16
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
100 d2q49a1
not modelled
99.5
16
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
101 d1pqua1
not modelled
99.4
17
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
102 d1mb4a1
not modelled
99.4
15
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
103 d1t4ba1
not modelled
99.4
21
Fold: NAD(P)-binding Rossmann-fold domainsSuperfamily: NAD(P)-binding Rossmann-fold domainsFamily: Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain
104 d2cvoa2
not modelled
99.3
11
Fold: FwdE/GAPDH domain-likeSuperfamily: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domainFamily: GAPDH-like
105 c3upyB_
not modelled
98.7
16
PDB header: oxidoreductaseChain: B: PDB Molecule: oxidoreductase;PDBTitle: crystal structure of the brucella abortus enzyme catalyzing the first2 committed step of the methylerythritol 4-phosphate pathway.
106 c5z2fA_
not modelled
98.6
18
PDB header: oxidoreductaseChain: A: PDB Molecule: dihydrodipicolinate reductase;PDBTitle: nadph/pda bound dihydrodipicolinate reductase from paenisporosarcina2 sp. tg-14
107 c5kt0A_
not modelled
98.6
17
PDB header: oxidoreductaseChain: A: PDB Molecule: 4-hydroxy-tetrahydrodipicolinate reductase;PDBTitle: dihydrodipicolinate reductase from the industrial and evolutionarily2 important cyanobacteria anabaena variabilis.
108 c4jn6B_
not modelled
98.5
16
PDB header: lyase/oxidoreductaseChain: B: PDB Molecule: acetaldehyde dehydrogenase;PDBTitle: crystal structure of the aldolase-dehydrogenase complex from2 mycobacterium tuberculosis hrv37
109 c4xb1B_
not modelled
98.4
18
PDB header: oxidoreductaseChain: B: PDB Molecule: 319aa long hypothetical homoserine dehydrogenase;PDBTitle: hyperthermophilic archaeal homoserine dehydrogenase in complex with2 nadph
110 c3euwB_
not modelled
98.3
22
PDB header: oxidoreductaseChain: B: PDB Molecule: myo-inositol dehydrogenase;PDBTitle: crystal structure of a myo-inositol dehydrogenase from corynebacterium2 glutamicum atcc 13032
111 c5ugjC_
not modelled
98.3
14
PDB header: oxidoreductaseChain: C: PDB Molecule: 4-hydroxy-tetrahydrodipicolinate reductase;PDBTitle: crystal structure of htpa reductase from neisseria meningitidis
112 c1nvmB_
not modelled
98.3
18
PDB header: lyase/oxidoreductaseChain: B: PDB Molecule: acetaldehyde dehydrogenase (acylating);PDBTitle: crystal structure of a bifunctional aldolase-dehydrogenase :2 sequestering a reactive and volatile intermediate
113 c3wgzB_
not modelled
98.2
24
PDB header: oxidoreductaseChain: B: PDB Molecule: meso-diaminopimelate dehydrogenase;PDBTitle: crystal structure of meso-dapdh q154l/t173i/r199m/p248s/h249n/n276s2 mutant with d-leucine of from clostridium tetani e88
114 c3wycB_
not modelled
98.2
26
PDB header: oxidoreductaseChain: B: PDB Molecule: meso-diaminopimelate d-dehydrogenase;PDBTitle: structure of a meso-diaminopimelate dehydrogenase in complex with nadp
115 c1drwA_
not modelled
98.2
28
PDB header: oxidoreductaseChain: A: PDB Molecule: dihydrodipicolinate reductase;PDBTitle: escherichia coli dhpr/nhdh complex
116 c3wb9A_
not modelled
98.2
22
PDB header: oxidoreductaseChain: A: PDB Molecule: diaminopimelate dehydrogenase;PDBTitle: crystal structures of meso-diaminopimelate dehydrogenase from2 symbiobacterium thermophilum
117 c5tenH_
not modelled
98.2
24
PDB header: oxidoreductaseChain: H: PDB Molecule: 4-hydroxy-tetrahydrodipicolinate reductase;PDBTitle: structure of 4-hydroxy-tetrahydrodipicolinate reductase from vibrio2 vulnificus with 2,5 furan dicarboxylic and nadh with intact3 polyhistidine tag
118 c4f3yA_
not modelled
98.1
25
PDB header: oxidoreductaseChain: A: PDB Molecule: dihydrodipicolinate reductase;PDBTitle: x-ray crystal structure of dihydrodipicolinate reductase from2 burkholderia thailandensis
119 c3ijpA_
not modelled
98.1
17
PDB header: oxidoreductaseChain: A: PDB Molecule: dihydrodipicolinate reductase;PDBTitle: crystal structure of dihydrodipicolinate reductase from bartonella2 henselae at 2.0a resolution
120 c3fd8A_
not modelled
98.1
27
PDB header: oxidoreductaseChain: A: PDB Molecule: oxidoreductase, gfo/idh/moca family;PDBTitle: crystal structure of an oxidoreductase from enterococcus2 faecalis