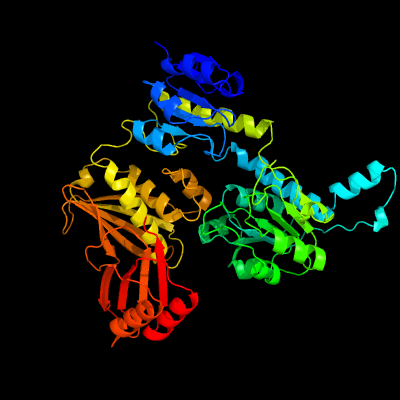
| 1 | c1ygyA_
|
|
 |
100.0 |
100 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-3-phosphoglycerate dehydrogenase;
PDBTitle: crystal structure of d-3-phosphoglycerate dehydrogenase from2 mycobacterium tuberculosis
|
|
|
|
| 2 | c1ybaC_
|
|
 |
100.0 |
34 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:d-3-phosphoglycerate dehydrogenase;
PDBTitle: the active form of phosphoglycerate dehydrogenase
|
|
|
|
| 3 | c3k5pA_
|
|
 |
100.0 |
31 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-3-phosphoglycerate dehydrogenase;
PDBTitle: crystal structure of amino acid-binding act: d-isomer specific 2-2 hydroxyacid dehydrogenase catalytic domain from brucella melitensis
|
|
|
|
| 4 | c4e5kC_
|
|
 |
100.0 |
32 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:phosphite dehydrogenase (thermostable variant);
PDBTitle: thermostable phosphite dehydrogenase in complex with nad and sulfite
|
|
|
|
| 5 | c4g2nA_
|
|
 |
100.0 |
28 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-isomer specific 2-hydroxyacid dehydrogenase, nad-binding;
PDBTitle: crystal structure of putative d-isomer specific 2-hydroxyacid2 dehydrogenase, nad-binding from polaromonas sp. js6 66
|
|
|
|
| 6 | c1gdhA_
|
|
 |
100.0 |
29 |
PDB header:oxidoreductase(choh (d)-nad(p)+ (a))
Chain: A: PDB Molecule:d-glycerate dehydrogenase;
PDBTitle: crystal structure of a nad-dependent d-glycerate2 dehydrogenase at 2.4 angstroms resolution
|
|
|
|
| 7 | c2gcgB_
|
|
 |
100.0 |
32 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:glyoxylate reductase/hydroxypyruvate reductase;
PDBTitle: ternary crystal structure of human glyoxylate2 reductase/hydroxypyruvate reductase
|
|
|
|
| 8 | c2d0iC_
|
|
 |
100.0 |
30 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:dehydrogenase;
PDBTitle: crystal structure ph0520 protein from pyrococcus horikoshii ot3
|
|
|
|
| 9 | c6ih2B_
|
|
 |
100.0 |
32 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:phosphite dehydrogenase;
PDBTitle: crystal structure of phosphite dehydrogenase from ralstonia sp. 4506
|
|
|
|
| 10 | c2dbqA_
|
|
 |
100.0 |
38 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:glyoxylate reductase;
PDBTitle: crystal structure of glyoxylate reductase (ph0597) from pyrococcus2 horikoshii ot3, complexed with nadp (i41)
|
|
|
|
| 11 | c2g76A_
|
|
 |
100.0 |
45 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-3-phosphoglycerate dehydrogenase;
PDBTitle: crystal structure of human 3-phosphoglycerate dehydrogenase
|
|
|
|
| 12 | c4lswA_
|
|
 |
100.0 |
26 |
PDB header:hydrolase
Chain: A: PDB Molecule:d-2-hydroxyacid dehydrogensase protein;
PDBTitle: crystallization and structural analysis of 2-hydroxyacid dehydrogenase2 from ketogulonicigenium vulgare y25
|
|
|
|
| 13 | c3n7uD_
|
|
 |
100.0 |
28 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:formate dehydrogenase;
PDBTitle: nad-dependent formate dehydrogenase from higher-plant arabidopsis2 thaliana in complex with nad and azide
|
|
|
|
| 14 | c3wnvA_
|
|
 |
100.0 |
32 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:glyoxylate reductase;
PDBTitle: crystal structure of a glyoxylate reductase from paecilomyes2 thermophila
|
|
|
|
| 15 | c5tx7A_
|
|
 |
100.0 |
34 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-isomer specific 2-hydroxyacid dehydrogenase family
PDBTitle: crystal structure of d-isomer specific 2-hydroxyacid dehydrogenase2 from desulfovibrio vulgaris
|
|
|
|
| 16 | c2eklA_
|
|
 |
100.0 |
35 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-3-phosphoglycerate dehydrogenase;
PDBTitle: structure of st1218 protein from sulfolobus tokodaii
|
|
|
|
| 17 | c3bazA_
|
|
 |
100.0 |
32 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:hydroxyphenylpyruvate reductase;
PDBTitle: structure of hydroxyphenylpyruvate reductase from coleus blumei in2 complex with nadp+
|
|
|
|
| 18 | c3fn4A_
|
|
 |
100.0 |
27 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:nad-dependent formate dehydrogenase;
PDBTitle: apo-form of nad-dependent formate dehydrogenase from bacterium2 moraxella sp.c-1 in closed conformation
|
|
|
|
| 19 | c3wwyA_
|
|
 |
100.0 |
27 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-lactate dehydrogenase;
PDBTitle: the crystal structure of d-lactate dehydrogenase from fusobacterium2 nucleatum subsp. nucleatum
|
|
|
|
| 20 | c4xkjA_
|
|
 |
100.0 |
27 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-lactate dehydrogenase;
PDBTitle: a novel d-lactate dehydrogenase from sporolactobacillus sp
|
|
|
|
| 21 | c2yq4C_ |
|
not modelled |
100.0 |
26 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:d-isomer specific 2-hydroxyacid dehydrogenase;
PDBTitle: crystal structure of d-isomer specific 2-hydroxyacid dehydrogenase2 from lactobacillus delbrueckii ssp. bulgaricus
|
|
|
| 22 | c5j23D_ |
|
not modelled |
100.0 |
29 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:2-hydroxyacid dehydrogenase;
PDBTitle: crystal structure of nadph-dependent glyoxylate/hydroxypyruvate2 reductase smc04462 (smghrb) from sinorhizobium meliloti in complex3 with 2'-phospho-adp-ribose
|
|
|
| 23 | c1j4aA_ |
|
not modelled |
100.0 |
31 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-lactate dehydrogenase;
PDBTitle: insights into domain closure, substrate specificity and2 catalysis of d-lactate dehydrogenase from lactobacillus3 bulgaricus
|
|
|
| 24 | c2omeA_ |
|
not modelled |
100.0 |
32 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:c-terminal-binding protein 2;
PDBTitle: crystal structure of human ctbp2 dehydrogenase complexed with nad(h)
|
|
|
| 25 | c4zgsE_ |
|
not modelled |
100.0 |
33 |
PDB header:oxidoreductase
Chain: E: PDB Molecule:putative d-lactate dehydrogenase;
PDBTitle: identification of the pyruvate reductase of chlamydomonas reinhardtii
|
|
|
| 26 | c3gg9C_ |
|
not modelled |
100.0 |
32 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:d-3-phosphoglycerate dehydrogenase oxidoreductase protein;
PDBTitle: crystal structure of putative d-3-phosphoglycerate dehydrogenase2 oxidoreductase from ralstonia solanacearum
|
|
|
| 27 | c1wwkA_ |
|
not modelled |
100.0 |
43 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:phosphoglycerate dehydrogenase;
PDBTitle: crystal structure of phosphoglycerate dehydrogenase from pyrococcus2 horikoshii ot3
|
|
|
| 28 | c3wwzB_ |
|
not modelled |
100.0 |
33 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:d-lactate dehydrogenase (fermentative);
PDBTitle: the crystal structure of d-lactate dehydrogenase from pseudomonas2 aeruginosa
|
|
|
| 29 | c2cukC_ |
|
not modelled |
100.0 |
39 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:glycerate dehydrogenase/glyoxylate reductase;
PDBTitle: crystal structure of tt0316 protein from thermus thermophilus hb8
|
|
|
| 30 | c4cukA_ |
|
not modelled |
100.0 |
29 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-lactate dehydrogenase;
PDBTitle: structure of salmonella d-lactate dehydrogenase in complex2 with nadh
|
|
|
| 31 | c2w2kB_ |
|
not modelled |
100.0 |
27 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:d-mandelate dehydrogenase;
PDBTitle: crystal structure of the apo forms of rhodotorula graminis2 d-mandelate dehydrogenase at 1.8a.
|
|
|
| 32 | c2nacA_ |
|
not modelled |
100.0 |
27 |
PDB header:oxidoreductase(aldehyde(d),nad+(a))
Chain: A: PDB Molecule:nad-dependent formate dehydrogenase;
PDBTitle: high resolution structures of holo and apo formate dehydrogenase
|
|
|
| 33 | c4prkB_ |
|
not modelled |
100.0 |
30 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:4-phosphoerythronate dehydrogenase;
PDBTitle: crystal structure of d-lactate dehydrogenase (d-ldh) from2 lactobacillus jensenii
|
|
|
| 34 | c1dxyA_ |
|
not modelled |
100.0 |
26 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-2-hydroxyisocaproate dehydrogenase;
PDBTitle: structure of d-2-hydroxyisocaproate dehydrogenase
|
|
|
| 35 | c2j6iC_ |
|
not modelled |
100.0 |
29 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:formate dehydrogenase;
PDBTitle: candida boidinii formate dehydrogenase (fdh) c-terminal mutant
|
|
|
| 36 | c6p2iA_ |
|
not modelled |
100.0 |
28 |
PDB header:oxidoreductase, biosynthetic protein
Chain: A: PDB Molecule:glycerate dehydrogenase;
PDBTitle: acyclic imino acid reductase (bsp5) in complex with nadph and d-arg
|
|
|
| 37 | c2pi1C_ |
|
not modelled |
100.0 |
28 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:d-lactate dehydrogenase;
PDBTitle: crystal structure of d-lactate dehydrogenase from aquifex2 aeolicus complexed with nad and lactic acid
|
|
|
| 38 | c1xdwA_ |
|
not modelled |
100.0 |
26 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:nad+-dependent (r)-2-hydroxyglutarate
PDBTitle: nad+-dependent (r)-2-hydroxyglutarate dehydrogenase from2 acidaminococcus fermentans
|
|
|
| 39 | c4s1vD_ |
|
not modelled |
100.0 |
28 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:d-3-phosphoglycerate dehydrogenase-related protein;
PDBTitle: crystal structure of phosphoglycerate oxidoreductase from vibrio2 cholerae o395
|
|
|
| 40 | c4xcvA_ |
|
not modelled |
100.0 |
28 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:nadp-dependent 2-hydroxyacid dehydrogenase;
PDBTitle: probable 2-hydroxyacid dehydrogenase from rhizobium etli cfn 42 in2 complex with nadph
|
|
|
| 41 | c3evtA_ |
|
not modelled |
100.0 |
25 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:phosphoglycerate dehydrogenase;
PDBTitle: crystal structure of phosphoglycerate dehydrogenase from lactobacillus2 plantarum
|
|
|
| 42 | c4zqbB_ |
|
not modelled |
100.0 |
28 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:nadp-dependent dehydrogenase;
PDBTitle: crystal structure of nadp-dependent dehydrogenase from2 rhodobactersphaeroides in complex with nadp and sulfate
|
|
|
| 43 | c3hg7A_ |
|
not modelled |
100.0 |
26 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-isomer specific 2-hydroxyacid dehydrogenase family
PDBTitle: crystal structure of d-isomer specific 2-hydroxyacid dehydrogenase2 family protein from aeromonas salmonicida subsp. salmonicida a449
|
|
|
| 44 | c4njmA_ |
|
not modelled |
100.0 |
28 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-3-phosphoglycerate dehydrogenase, putative;
PDBTitle: crystal structure of phosphoglycerate bound 3-phosphoglycerate2 dehydrogenase in entamoeba histolytica
|
|
|
| 45 | c4n18A_ |
|
not modelled |
100.0 |
24 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-isomer specific 2-hydroxyacid dehydrogenase family
PDBTitle: crystal structure of d-isomer specific 2-hydroxyacid dehydrogenase2 family protein from klebsiella pneumoniae 342
|
|
|
| 46 | c1qp8A_ |
|
not modelled |
100.0 |
25 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:formate dehydrogenase;
PDBTitle: crystal structure of a putative formate dehydrogenase from2 pyrobaculum aerophilum
|
|
|
| 47 | c5mh5A_ |
|
not modelled |
100.0 |
25 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-2-hydroxyacid dehydrogenase;
PDBTitle: d-2-hydroxyacid dehydrogenases (d2-hdh) from haloferax mediterranei in2 complex with 2-keto-hexanoic acid and nadp+ (1.4 a resolution)
|
|
|
| 48 | c4weqA_ |
|
not modelled |
100.0 |
27 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:nad-dependent dehydrogenase;
PDBTitle: crystal structure of nadph-dependent glyoxylate/hydroxypyruvate2 reductase smc02828 (smghra) from sinorhizobium meliloti in complex3 with nadp and sulfate
|
|
|
| 49 | c4hy3D_ |
|
not modelled |
100.0 |
26 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:phosphoglycerate oxidoreductase;
PDBTitle: crystal structure of a phosphoglycerate oxidoreductase from rhizobium2 etli
|
|
|
| 50 | c5dt9A_ |
|
not modelled |
100.0 |
33 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:erythronate-4-phosphate dehydrogenase;
PDBTitle: crystal structure of a putative d-erythronate-4-phosphate2 dehydrogenase from vibrio cholerae
|
|
|
| 51 | c2o4cB_ |
|
not modelled |
100.0 |
32 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:erythronate-4-phosphate dehydrogenase;
PDBTitle: crystal structure of d-erythronate-4-phosphate dehydrogenase complexed2 with nad
|
|
|
| 52 | c3kboB_ |
|
not modelled |
100.0 |
26 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:glyoxylate/hydroxypyruvate reductase a;
PDBTitle: 2.14 angstrom crystal structure of putative oxidoreductase (ycdw) from2 salmonella typhimurium in complex with nadp
|
|
|
| 53 | c3oetF_ |
|
not modelled |
100.0 |
28 |
PDB header:oxidoreductase
Chain: F: PDB Molecule:erythronate-4-phosphate dehydrogenase;
PDBTitle: d-erythronate-4-phosphate dehydrogenase complexed with nad
|
|
|
| 54 | c4xa8A_ |
|
not modelled |
100.0 |
30 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:d-isomer specific 2-hydroxyacid dehydrogenase nad-binding;
PDBTitle: crystal structure of d-isomer specific 2-hydroxyacid dehydrogenase2 from xanthobacter autotrophicus py2
|
|
|
| 55 | c4dgsA_ |
|
not modelled |
100.0 |
33 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:dehydrogenase;
PDBTitle: the crystals structure of dehydrogenase from rhizobium meliloti
|
|
|
| 56 | c3gvxA_ |
|
not modelled |
100.0 |
26 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:glycerate dehydrogenase related protein;
PDBTitle: crystal structure of glycerate dehydrogenase related2 protein from thermoplasma acidophilum
|
|
|
| 57 | d1ygya1 |
|
not modelled |
100.0 |
100 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 58 | d1gdha1 |
|
not modelled |
100.0 |
32 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 59 | d1mx3a1 |
|
not modelled |
100.0 |
38 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 60 | d1j4aa1 |
|
not modelled |
100.0 |
36 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 61 | d2dlda1 |
|
not modelled |
100.0 |
35 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 62 | d1dxya1 |
|
not modelled |
100.0 |
32 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 63 | d2naca1 |
|
not modelled |
100.0 |
29 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 64 | d1sc6a1 |
|
not modelled |
100.0 |
34 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 65 | d1qp8a1 |
|
not modelled |
100.0 |
28 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 66 | c5uscB_ |
|
not modelled |
100.0 |
17 |
PDB header:hydrolase
Chain: B: PDB Molecule:prephenate dehydrogenase;
PDBTitle: crystal structure of prephenate dehydrogenase tyra from bacillus2 anthracis in complex with nad and l-tyrosine
|
|
|
| 67 | c1l7eC_ |
|
not modelled |
100.0 |
22 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:nicotinamide nucleotide transhydrogenase,
PDBTitle: crystal structure of r. rubrum transhydrogenase domain i2 with bound nadh
|
|
|
| 68 | d1ygya4 |
|
not modelled |
100.0 |
100 |
Fold:FwdE/GAPDH domain-like
Superfamily:Serine metabolism enzymes domain
Family:SerA intervening domain-like |
|
|
| 69 | c5v96A_ |
|
not modelled |
100.0 |
17 |
PDB header:hydrolase
Chain: A: PDB Molecule:s-adenosyl-l-homocysteine hydrolase;
PDBTitle: crystal structure of s-adenosyl-l-homocysteine hydrolase from2 naegleria fowleri with bound nad and adenosine
|
|
|
| 70 | c6f3oC_ |
|
not modelled |
100.0 |
16 |
PDB header:hydrolase
Chain: C: PDB Molecule:adenosylhomocysteinase;
PDBTitle: crystal structure of s-adenosyl-l-homocysteine hydrolase from2 pseudomonas aeruginosa complexed with adenine, k+ and zn2+ cations
|
|
|
| 71 | c1d4fD_ |
|
not modelled |
100.0 |
18 |
PDB header:hydrolase
Chain: D: PDB Molecule:s-adenosylhomocysteine hydrolase;
PDBTitle: crystal structure of recombinant rat-liver d244e mutant s-2 adenosylhomocysteine hydrolase
|
|
|
| 72 | c3gvpB_ |
|
not modelled |
100.0 |
16 |
PDB header:hydrolase
Chain: B: PDB Molecule:adenosylhomocysteinase 3;
PDBTitle: human sahh-like domain of human adenosylhomocysteinase 3
|
|
|
| 73 | c5hm8C_ |
|
not modelled |
100.0 |
16 |
PDB header:hydrolase
Chain: C: PDB Molecule:adenosylhomocysteinase;
PDBTitle: 2.85 angstrom crystal structure of s-adenosylhomocysteinase from2 cryptosporidium parvum in complex with adenosine and nad.
|
|
|
| 74 | c6aphA_ |
|
not modelled |
100.0 |
18 |
PDB header:hydrolase
Chain: A: PDB Molecule:adenosylhomocysteinase;
PDBTitle: crystal structure of adenosylhomocysteinase from elizabethkingia2 anophelis nuhp1 in complex with nad and adenosine
|
|
|
| 75 | c3dhyC_ |
|
not modelled |
100.0 |
19 |
PDB header:hydrolase
Chain: C: PDB Molecule:adenosylhomocysteinase;
PDBTitle: crystal structures of mycobacterium tuberculosis s-adenosyl-l-2 homocysteine hydrolase in ternary complex with substrate and3 inhibitors
|
|
|
| 76 | c3n58D_ |
|
not modelled |
100.0 |
21 |
PDB header:hydrolase
Chain: D: PDB Molecule:adenosylhomocysteinase;
PDBTitle: crystal structure of s-adenosyl-l-homocysteine hydrolase from brucella2 melitensis in ternary complex with nad and adenosine, orthorhombic3 form
|
|
|
| 77 | c1v8bA_ |
|
not modelled |
100.0 |
18 |
PDB header:hydrolase
Chain: A: PDB Molecule:adenosylhomocysteinase;
PDBTitle: crystal structure of a hydrolase
|
|
|
| 78 | c3oneA_ |
|
not modelled |
100.0 |
17 |
PDB header:hydrolase/hydrolase substrate
Chain: A: PDB Molecule:adenosylhomocysteinase;
PDBTitle: crystal structure of lupinus luteus s-adenosyl-l-homocysteine2 hydrolase in complex with adenine
|
|
|
| 79 | c3x2fA_ |
|
not modelled |
100.0 |
17 |
PDB header:hydrolase
Chain: A: PDB Molecule:adenosylhomocysteinase;
PDBTitle: a thermophilic s-adenosylhomocysteine hydrolase
|
|
|
| 80 | c2bruB_ |
|
not modelled |
100.0 |
21 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:nad(p) transhydrogenase subunit alpha;
PDBTitle: complex of the domain i and domain iii of escherichia coli2 transhydrogenase
|
|
|
| 81 | c1pjcA_ |
|
not modelled |
100.0 |
18 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:protein (l-alanine dehydrogenase);
PDBTitle: l-alanine dehydrogenase complexed with nad
|
|
|
| 82 | c2eezG_ |
|
not modelled |
100.0 |
17 |
PDB header:oxidoreductase
Chain: G: PDB Molecule:alanine dehydrogenase;
PDBTitle: crystal structure of alanine dehydrogenase from themus thermophilus
|
|
|
| 83 | c2vhyB_ |
|
not modelled |
100.0 |
17 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:alanine dehydrogenase;
PDBTitle: crystal structure of apo l-alanine dehydrogenase from mycobacterium2 tuberculosis
|
|
|
| 84 | d1ygya2 |
|
not modelled |
100.0 |
79 |
Fold:Flavodoxin-like
Superfamily:Formate/glycerate dehydrogenase catalytic domain-like
Family:Formate/glycerate dehydrogenases, substrate-binding domain |
|
|
| 85 | c4izhA_ |
|
not modelled |
100.0 |
24 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:nad/nadp transhydrogenase alpha subunit 1;
PDBTitle: crystal structure of the alpha1 dimer of thermus thermophilus2 transhydrogenase in p6
|
|
|
| 86 | c3p2yA_ |
|
not modelled |
99.9 |
18 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:alanine dehydrogenase/pyridine nucleotide transhydrogenase;
PDBTitle: crystal structure of alanine dehydrogenase/pyridine nucleotide2 transhydrogenase from mycobacterium smegmatis
|
|
|
| 87 | c3d64A_ |
|
not modelled |
99.9 |
19 |
PDB header:hydrolase
Chain: A: PDB Molecule:adenosylhomocysteinase;
PDBTitle: crystal structure of s-adenosyl-l-homocysteine hydrolase from2 burkholderia pseudomallei
|
|
|
| 88 | d1v8ba1 |
|
not modelled |
99.9 |
22 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 89 | d1sc6a2 |
|
not modelled |
99.9 |
23 |
Fold:Flavodoxin-like
Superfamily:Formate/glycerate dehydrogenase catalytic domain-like
Family:Formate/glycerate dehydrogenases, substrate-binding domain |
|
|
| 90 | c4dioB_ |
|
not modelled |
99.9 |
19 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:nad(p) transhydrogenase subunit alpha part 1;
PDBTitle: the crystal structure of transhydrogenase from sinorhizobium meliloti
|
|
|
| 91 | d1gdha2 |
|
not modelled |
99.9 |
25 |
Fold:Flavodoxin-like
Superfamily:Formate/glycerate dehydrogenase catalytic domain-like
Family:Formate/glycerate dehydrogenases, substrate-binding domain |
|
|
| 92 | d1np3a2 |
|
not modelled |
99.9 |
22 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
|
|
| 93 | d1leha1 |
|
not modelled |
99.9 |
14 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Aminoacid dehydrogenase-like, C-terminal domain |
|
|
| 94 | d1li4a1 |
|
not modelled |
99.9 |
20 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 95 | d1mx3a2 |
|
not modelled |
99.9 |
27 |
Fold:Flavodoxin-like
Superfamily:Formate/glycerate dehydrogenase catalytic domain-like
Family:Formate/glycerate dehydrogenases, substrate-binding domain |
|
|
| 96 | d2naca2 |
|
not modelled |
99.9 |
22 |
Fold:Flavodoxin-like
Superfamily:Formate/glycerate dehydrogenase catalytic domain-like
Family:Formate/glycerate dehydrogenases, substrate-binding domain |
|
|
| 97 | d1dxya2 |
|
not modelled |
99.9 |
15 |
Fold:Flavodoxin-like
Superfamily:Formate/glycerate dehydrogenase catalytic domain-like
Family:Formate/glycerate dehydrogenases, substrate-binding domain |
|
|
| 98 | c3d4oA_ |
|
not modelled |
99.8 |
19 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:dipicolinate synthase subunit a;
PDBTitle: crystal structure of dipicolinate synthase subunit a (np_243269.1)2 from bacillus halodurans at 2.10 a resolution
|
|
|
| 99 | d1pjca1 |
|
not modelled |
99.8 |
23 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 100 | c2rirA_ |
|
not modelled |
99.8 |
18 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:dipicolinate synthase, a chain;
PDBTitle: crystal structure of dipicolinate synthase, a chain, from bacillus2 subtilis
|
|
|
| 101 | d1ygya3 |
|
not modelled |
99.8 |
100 |
Fold:Ferredoxin-like
Superfamily:ACT-like
Family:Phosphoglycerate dehydrogenase, regulatory (C-terminal) domain |
|
|
| 102 | d1j4aa2 |
|
not modelled |
99.8 |
22 |
Fold:Flavodoxin-like
Superfamily:Formate/glycerate dehydrogenase catalytic domain-like
Family:Formate/glycerate dehydrogenases, substrate-binding domain |
|
|
| 103 | d2dlda2 |
|
not modelled |
99.8 |
17 |
Fold:Flavodoxin-like
Superfamily:Formate/glycerate dehydrogenase catalytic domain-like
Family:Formate/glycerate dehydrogenases, substrate-binding domain |
|
|
| 104 | d2iafa1 |
|
not modelled |
99.7 |
9 |
Fold:FwdE/GAPDH domain-like
Superfamily:Serine metabolism enzymes domain
Family:Serine dehydratase beta chain-like |
|
|
| 105 | c5b37A_ |
|
not modelled |
99.7 |
15 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:tryptophan dehydrogenase;
PDBTitle: crystal structure of l-tryptophan dehydrogenase from nostoc2 punctiforme
|
|
|
| 106 | d1sc6a3 |
|
not modelled |
99.6 |
25 |
Fold:Ferredoxin-like
Superfamily:ACT-like
Family:Phosphoglycerate dehydrogenase, regulatory (C-terminal) domain |
|
|
| 107 | d1c1da1 |
|
not modelled |
99.6 |
24 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Aminoacid dehydrogenase-like, C-terminal domain |
|
|
| 108 | c1lehB_ |
|
not modelled |
99.6 |
18 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:leucine dehydrogenase;
PDBTitle: leucine dehydrogenase from bacillus sphaericus
|
|
|
| 109 | d1l7da1 |
|
not modelled |
99.5 |
24 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
|
|
| 110 | c4rqoB_ |
|
not modelled |
99.5 |
13 |
PDB header:lyase
Chain: B: PDB Molecule:l-serine dehydratase;
PDBTitle: crystal structure of l-serine dehydratase from legionella pneumophila
|
|
|
| 111 | c6jczL_ |
|
not modelled |
99.4 |
16 |
PDB header:isomerase
Chain: L: PDB Molecule:putative ketol-acid reductoisomerase 2;
PDBTitle: cryo-em structure of sulfolobus solfataricus ketol-acid2 reductoisomerase (sso-kari) in complex with mg2+, nadph, and cpd at3 ph7.5
|
|
|
| 112 | c4tskA_ |
|
not modelled |
99.4 |
21 |
PDB header:oxidoreductase,isomerase
Chain: A: PDB Molecule:ketol-acid reductoisomerase;
PDBTitle: ketol-acid reductoisomerase from alicyclobacillus acidocaldarius
|
|
|
| 113 | c6aqjB_ |
|
not modelled |
99.4 |
22 |
PDB header:isomerase
Chain: B: PDB Molecule:ketol-acid reductoisomerase (nadp(+));
PDBTitle: crystal structures of staphylococcus aureus ketol-acid2 reductoisomerase in complex with two transition state analogs that3 have biocidal activity.
|
|
|
| 114 | c4ypoB_ |
|
not modelled |
99.3 |
23 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:ketol-acid reductoisomerase;
PDBTitle: crystal structure of mycobacterium tuberculosis ketol-acid2 reductoisomerase in complex with mg2+
|
|
|
| 115 | c4kqxB_ |
|
not modelled |
99.2 |
26 |
PDB header:oxidoreductase/oxidoreductase inhibitor
Chain: B: PDB Molecule:ketol-acid reductoisomerase;
PDBTitle: mutant slackia exigua kari ddv in complex with nad and an inhibitor
|
|
|
| 116 | c5yeqB_ |
|
not modelled |
99.2 |
24 |
PDB header:isomerase
Chain: B: PDB Molecule:ketol-acid reductoisomerase (nadp(+));
PDBTitle: the structure of sac-kari protein
|
|
|
| 117 | c4xdzB_ |
|
not modelled |
99.2 |
25 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:ketol-acid reductoisomerase;
PDBTitle: holo structure of ketol-acid reductoisomerase from ignisphaera2 aggregans
|
|
|
| 118 | c1np3B_ |
|
not modelled |
99.2 |
24 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:ketol-acid reductoisomerase;
PDBTitle: crystal structure of class i acetohydroxy acid isomeroreductase from2 pseudomonas aeruginosa
|
|
|
| 119 | c3oj0A_ |
|
not modelled |
99.2 |
16 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:glutamyl-trna reductase;
PDBTitle: crystal structure of glutamyl-trna reductase from thermoplasma2 volcanium (nucleotide binding domain)
|
|
|
| 120 | d3cuma2 |
|
not modelled |
99.2 |
20 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
|
|