1 c3hfeC_
54.1
28
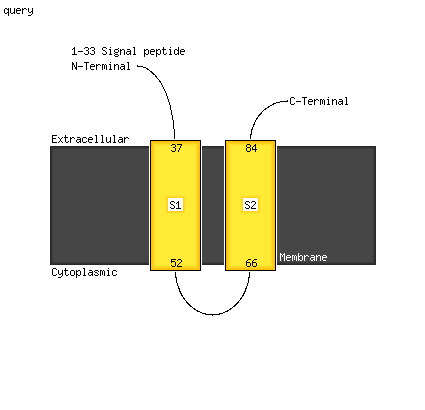
PDB header: transport proteinChain: C: PDB Molecule: potassium voltage-gated channel subfamily kqt member 1;PDBTitle: a trimeric form of the kv7.1 a domain tail
2 c3bj4B_
47.9
26
PDB header: signaling proteinChain: B: PDB Molecule: potassium voltage-gated channel subfamily kqtPDBTitle: the kcnq1 (kv7.1) c-terminus, a multi-tiered scaffold for2 subunit assembly and protein interaction
3 c3ra3D_
29.8
18
PDB header: de novo proteinChain: D: PDB Molecule: p2f;PDBTitle: crystal structure of a section of a de novo design gigadalton protein2 fibre
4 c5b16C_
25.6
20
PDB header: hydrolaseChain: C: PDB Molecule: microprocessor complex subunit dgcr8;PDBTitle: x-ray structure of drosha in complex with the c-terminal tail of2 dgcr8.
5 c5b16B_
24.7
20
PDB header: hydrolaseChain: B: PDB Molecule: microprocessor complex subunit dgcr8;PDBTitle: x-ray structure of drosha in complex with the c-terminal tail of2 dgcr8.
6 c3ra3B_
24.2
20
PDB header: de novo proteinChain: B: PDB Molecule: p2f;PDBTitle: crystal structure of a section of a de novo design gigadalton protein2 fibre
7 c1junB_
23.9
18
PDB header: transcription regulationChain: B: PDB Molecule: c-jun homodimer;PDBTitle: nmr study of c-jun homodimer
8 c1ce0B_
23.2
13
PDB header: hiv-1 envelope proteinChain: B: PDB Molecule: protein (leucine zipper model h38-p1);PDBTitle: trimerization specificity in hiv-1 gp41: analysis with a2 gcn4 leucine zipper model
9 c2jeeA_
20.0
20
PDB header: cell cycleChain: A: PDB Molecule: cell division protein zapb;PDBTitle: xray structure of e. coli yiiu
10 c1ij2C_
19.8
18
PDB header: transcriptionChain: C: PDB Molecule: general control protein gcn4;PDBTitle: gcn4-pvtl coiled-coil trimer with threonine at the a(16)2 position
11 c1ij3B_
19.3
18
PDB header: transcriptionChain: B: PDB Molecule: general control protein gcn4;PDBTitle: gcn4-pvsl coiled-coil trimer with serine at the a(16)2 position
12 c1ij3C_
19.3
18
PDB header: transcriptionChain: C: PDB Molecule: general control protein gcn4;PDBTitle: gcn4-pvsl coiled-coil trimer with serine at the a(16)2 position
13 c3k7zB_
19.2
18
PDB header: dna binding proteinChain: B: PDB Molecule: general control protein gcn4;PDBTitle: gcn4-leucine zipper core mutant as n16a trigonal automatic2 solution
14 c3k7zA_
19.2
18
PDB header: dna binding proteinChain: A: PDB Molecule: general control protein gcn4;PDBTitle: gcn4-leucine zipper core mutant as n16a trigonal automatic2 solution
15 c1rb6C_
19.2
18
PDB header: dna binding proteinChain: C: PDB Molecule: general control protein gcn4;PDBTitle: antiparallel trimer of gcn4-leucine zipper core mutant as n16a2 tetragonal form
16 c1rb1B_
19.2
18
PDB header: dna binding proteinChain: B: PDB Molecule: general control protein gcn4;PDBTitle: gcn4-leucine zipper core mutant as n16a trigonal automatic2 solution
17 c1swiA_
19.2
18
PDB header: leucine zipperChain: A: PDB Molecule: gcn4p1;PDBTitle: gcn4-leucine zipper core mutant as n16a complexed with benzene
18 c1rb1A_
19.2
18
PDB header: dna binding proteinChain: A: PDB Molecule: general control protein gcn4;PDBTitle: gcn4-leucine zipper core mutant as n16a trigonal automatic2 solution
19 c1zxaB_
18.1
22
PDB header: transferaseChain: B: PDB Molecule: cgmp-dependent protein kinase 1, alpha isozyme;PDBTitle: solution structure of the coiled-coil domain of cgmp-2 dependent protein kinase ia
20 c1ij2B_
17.9
18
PDB header: transcriptionChain: B: PDB Molecule: general control protein gcn4;PDBTitle: gcn4-pvtl coiled-coil trimer with threonine at the a(16)2 position
21 c2o7hF_
not modelled
17.8
18
PDB header: transcriptionChain: F: PDB Molecule: general control protein gcn4;PDBTitle: crystal structure of trimeric coiled coil gcn4 leucine zipper
22 c3mk7F_
not modelled
17.0
13
PDB header: oxidoreductaseChain: F: PDB Molecule: cytochrome c oxidase, cbb3-type, subunit p;PDBTitle: the structure of cbb3 cytochrome oxidase
23 c5yr0B_
not modelled
16.7
20
PDB header: endocytosisChain: B: PDB Molecule: uv radiation resistance associated protein;PDBTitle: structure of beclin1-uvrag coiled coil domain complex
24 c4r4mB_
not modelled
15.1
22
PDB header: dna binding proteinChain: B: PDB Molecule: cgmp-dependent protein kinase 1;PDBTitle: crystal structure of c42l cgmp dependent protein kinase i alpha (pkgi2 alpha) leucine zipper
25 c3vgxD_
not modelled
13.1
33
PDB header: membrane proteinChain: D: PDB Molecule: envelope glycoprotein gp160;PDBTitle: structure of gp41 t21/cp621-652
26 c1ztaA_
not modelled
12.7
17
PDB header: dna-binding motifChain: A: PDB Molecule: leucine zipper monomer;PDBTitle: the solution structure of a leucine-zipper motif peptide
27 c1u2uA_
not modelled
12.2
12
PDB header: transcriptionChain: A: PDB Molecule: general control protein gcn4;PDBTitle: nmr solution structure of a designed heterodimeric leucine2 zipper
28 c4u5tB_
not modelled
12.2
15
PDB header: transcription/transcription inhibitorChain: B: PDB Molecule: vbp leucine zipper;PDBTitle: crystal structure of vbp leucine zipper with bound arylstibonic acid
29 c2xzrA_
not modelled
12.0
13
PDB header: cell adhesionChain: A: PDB Molecule: immunoglobulin-binding protein eibd;PDBTitle: escherichia coli immunoglobulin-binding protein eibd 391-438 fused2 to gcn4 adaptors
30 c4yj0C_
not modelled
10.7
32
PDB header: transcriptionChain: C: PDB Molecule: doublesex- and mab-3-related transcription factor 1;PDBTitle: crystal structure of the dm domain of human dmrt1 bound to 25mer2 target dna
31 c3iynR_
not modelled
10.5
9
PDB header: virusChain: R: PDB Molecule: hexon-associated protein;PDBTitle: 3.6-angstrom cryoem structure of human adenovirus type 5
32 c1ci6B_
not modelled
10.1
15
PDB header: transcriptionChain: B: PDB Molecule: transcription factor c/ebp beta;PDBTitle: transcription factor atf4-c/ebp beta bzip heterodimer
33 c6mctJ_
not modelled
9.9
24
PDB header: de novo proteinChain: J: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
34 c6mctE_
not modelled
9.9
24
PDB header: de novo proteinChain: E: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
35 c6mctI_
not modelled
9.9
24
PDB header: de novo proteinChain: I: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
36 c6mctH_
not modelled
9.9
24
PDB header: de novo proteinChain: H: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
37 c6mq2D_
not modelled
9.9
24
PDB header: de novo proteinChain: D: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-2
38 c6mctD_
not modelled
9.9
24
PDB header: de novo proteinChain: D: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
39 c6mctK_
not modelled
9.9
24
PDB header: de novo proteinChain: K: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
40 c6mctM_
not modelled
9.9
24
PDB header: de novo proteinChain: M: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
41 c6mctF_
not modelled
9.9
24
PDB header: de novo proteinChain: F: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
42 c6mctB_
not modelled
9.9
24
PDB header: de novo proteinChain: B: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
43 c6mctN_
not modelled
9.9
24
PDB header: de novo proteinChain: N: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
44 c6mctL_
not modelled
9.9
24
PDB header: de novo proteinChain: L: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
45 c6mctA_
not modelled
9.9
24
PDB header: de novo proteinChain: A: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
46 c6mctO_
not modelled
9.9
24
PDB header: de novo proteinChain: O: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
47 c6mctC_
not modelled
9.9
24
PDB header: de novo proteinChain: C: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
48 c6mctG_
not modelled
9.9
24
PDB header: de novo proteinChain: G: PDB Molecule: mini-evgl membrane protein;PDBTitle: a designed pentameric membrane protein stabilized by van der waals2 interaction
49 c6mpwA_
not modelled
9.9
24
PDB header: de novo proteinChain: A: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-1
50 c6mpwD_
not modelled
9.7
24
PDB header: de novo proteinChain: D: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-1
51 c6mpwE_
not modelled
9.7
24
PDB header: de novo proteinChain: E: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-1
52 c6mq2A_
not modelled
9.7
24
PDB header: de novo proteinChain: A: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-2
53 c6mq2C_
not modelled
9.7
24
PDB header: de novo proteinChain: C: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-2
54 c6mpwB_
not modelled
9.7
24
PDB header: de novo proteinChain: B: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-1
55 c6mq2B_
not modelled
9.7
24
PDB header: de novo proteinChain: B: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-2
56 c6mpwC_
not modelled
9.7
24
PDB header: de novo proteinChain: C: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-1
57 c6mq2E_
not modelled
9.7
24
PDB header: de novo proteinChain: E: PDB Molecule: mini-evgl membrane protein;PDBTitle: de novo design of membrane protein--mini-evgl membrane protein, c22212 form-2
58 c4px7A_
not modelled
9.6
11
PDB header: hydrolaseChain: A: PDB Molecule: phosphatidylglycerophosphatase;PDBTitle: crystal structure of lipid phosphatase e. coli pgpb
59 c5j10A_
not modelled
9.2
13
PDB header: de novo proteinChain: A: PDB Molecule: peptide design 2l4hc2_24;PDBTitle: de novo design of protein homo-oligomers with modular hydrogen bond2 network-mediated specificity
60 c1dipA_
not modelled
8.8
15
PDB header: acetylationChain: A: PDB Molecule: delta-sleep-inducing peptide immunoreactivePDBTitle: the solution structure of porcine delta-sleep-inducing2 peptide immunoreactive peptide, nmr, 10 structures
61 c2kluA_
not modelled
8.4
14
PDB header: immune system, membrane proteinChain: A: PDB Molecule: t-cell surface glycoprotein cd4;PDBTitle: nmr structure of the transmembrane and cytoplasmic domains2 of human cd4
62 c1pl5A_
not modelled
8.3
13
PDB header: dna binding protein/transcriptionChain: A: PDB Molecule: regulatory protein sir4;PDBTitle: crystal structure analysis of the sir4p c-terminal coiled2 coil
63 c3iynQ_
not modelled
8.1
16
PDB header: virusChain: Q: PDB Molecule: hexon-associated protein;PDBTitle: 3.6-angstrom cryoem structure of human adenovirus type 5
64 c1nyhA_
not modelled
8.1
13
PDB header: transcription repressorChain: A: PDB Molecule: regulatory protein sir4;PDBTitle: crystal structure of the coiled-coil dimerization motif of sir4
65 c2na6C_
not modelled
6.7
23
PDB header: apoptosisChain: C: PDB Molecule: tumor necrosis factor receptor superfamily member 6;PDBTitle: transmembrane domain of mouse fas/cd95 death receptor
66 c2na6A_
not modelled
6.7
23
PDB header: apoptosisChain: A: PDB Molecule: tumor necrosis factor receptor superfamily member 6;PDBTitle: transmembrane domain of mouse fas/cd95 death receptor
67 c2na6B_
not modelled
6.7
23
PDB header: apoptosisChain: B: PDB Molecule: tumor necrosis factor receptor superfamily member 6;PDBTitle: transmembrane domain of mouse fas/cd95 death receptor
68 c5dolB_
not modelled
5.9
18
PDB header: replicationChain: B: PDB Molecule: initiation-control protein yaba;PDBTitle: crystal structure of yaba amino-terminal domain from bacillus subtilis
69 c2oqqB_
not modelled
5.5
13
PDB header: transcriptionChain: B: PDB Molecule: transcription factor hy5;PDBTitle: crystal structure of hy5 leucine zipper homodimer from arabidopsis2 thaliana