| H37Rv Coordinates: | 3817419 – 3818207 |
|---|---|
| Vulnerability Index (VI): | 1.0520 |
| M. smegmatis homologs | MSMEG1707 |
| M. abscessus homologs |
DNA Sequence
>rv3400_Rv3400_dnaTGCGCATGAAATCCGCGGCCGAGGCATGGCGGGAACGCGGCTTTGATCTGGACTTGACCGAGCTGATCTACTTCGATCAACGCAACGACGTGGCCGACTACCTCGCCGGCTCCGGCTGGCAGGTCACCACCAGCACCGGCAAGGAACTCTTTGCGGCCCAAGGGCTGCCGCCCTTCGCGGACGACCACATAACTCGGTTCGCCGACCGCCGCTACATCAGCGCGGTGCTGAAGTAGGTGGCCCCGGCACTATAGCCGGGCCTAACTCGTAGGCTTGGTACGCGGGCAGAGCCGCCAGGCATGGCGAACTGGTATCGCCCGAACTATCCGGAAGTGAGGTCCCGCGTGCTGGGTCTGCCCGAGAAGGTGCGTGCTTGCCTGTTCGACCTCGACGGTGTGCTCACCGATACCGCGAGCCTGCATACCAAGGCGTGGAAGGCCATGTTTGACGCCTACCTAGCCGAGCGAGCCGAGCGCACCGGCGAAAAATTCGTTCCCTTCGACCCTGCCGCGGACTATCACACGTATGTGGACGGCAAGAAACGCGAAGACGGCGTTCGATCGTTTCTGAGCAGCCGCGCCATCGAAATACCCGACGGTTCCCCGGATGACCCGGGCGCCGCCGAGACGGTGTATGGCCTGGGCAACCGCAAGAACGACATGTTGCACAAGCTGCTGCGCGACGATGGGGCCCAGGTGTTCGACGGGTCGCGGCGCTACCTGGAGGCGGTCACGGCCGCGGGTCTCGGTGTGGCCGTGGTGTCTTCGAGCGCCAACACCCGCGACGTGCTCGCGACCACCGGTCTGGACCGGTTCGTCCAGCAGCGGGTGGACGGCGTGACGTTGCGCGAAGAGCACATCGCCGGCAAGCCGGCCCCCGACTCCTTCCTGCGCGCGGCAGAACTGTTGGGGGTTACCCCCGACGCGGCGGCGGTGTTCGAGGACGCCCTGTCCGGGGTGGCGGCCGGCCGCGCCGGCAACTTCGCCGTAGTGGTGGGCATCAACCGAACGGGCCGGGCGGCTCAGGCCGCCCAGTTGCGCCGCCATGGCGCCGACGTGGTGGTAACCGATCTCGCCGAGCTGCTGTAG
Protein Sequence
>Rv3400_Rv3400_proteinMANWYRPNYPEVRSRVLGLPEKVRACLFDLDGVLTDTASLHTKAWKAMFDAYLAERAERTGEKFVPFDPAADYHTYVDGKKREDGVRSFLSSRAIEIPDGSPDDPGAAETVYGLGNRKNDMLHKLLRDDGAQVFDGSRRYLEAVTAAGLGVAVVSSSANTRDVLATTGLDRFVQQRVDGVTLREEHIAGKPAPDSFLRAAELLGVTPDAAAVFEDALSGVAAGRAGNFAVVVGINRTGRAAQAAQLRRHGADVVVTDLAELL
Essentiality:
| Call | Condition | Type | Source | Notes |
|---|---|---|---|---|
| Non Essential | in vitro | TnSeq | [DeJesus et al. (2017)] | |
| Non Essential | in vitro | CRISPRi | [Bosch & DeJesus et al. (2021)] |
Experiment (PMID: 34297925): We designed an M. tuberculosis CRISPRi library (RLC12) to target all annotated Mtb genes with single guide RNAs (sgRNAs) of varying predicted knockdown efficiencies.
In parallel, we constructed a M. smegmatis CRISPRi library (RLC11) to target all annotated Msmeg genes, similar to the Mtb sgRNA library.
RLC12 was transformed into M. tuberculosis H37Rv; RLC11 into M. smegmatis mc2-155.
We passaged both CRISPRi libraries for nearly 30 generations in the presence or absence of the CRISPRi-inducer anhydrotetracycline (ATc). At the indicated timepoints, genomic DNA was harvested and the representation of individual sgRNAs was counted using Illumina sequencing.
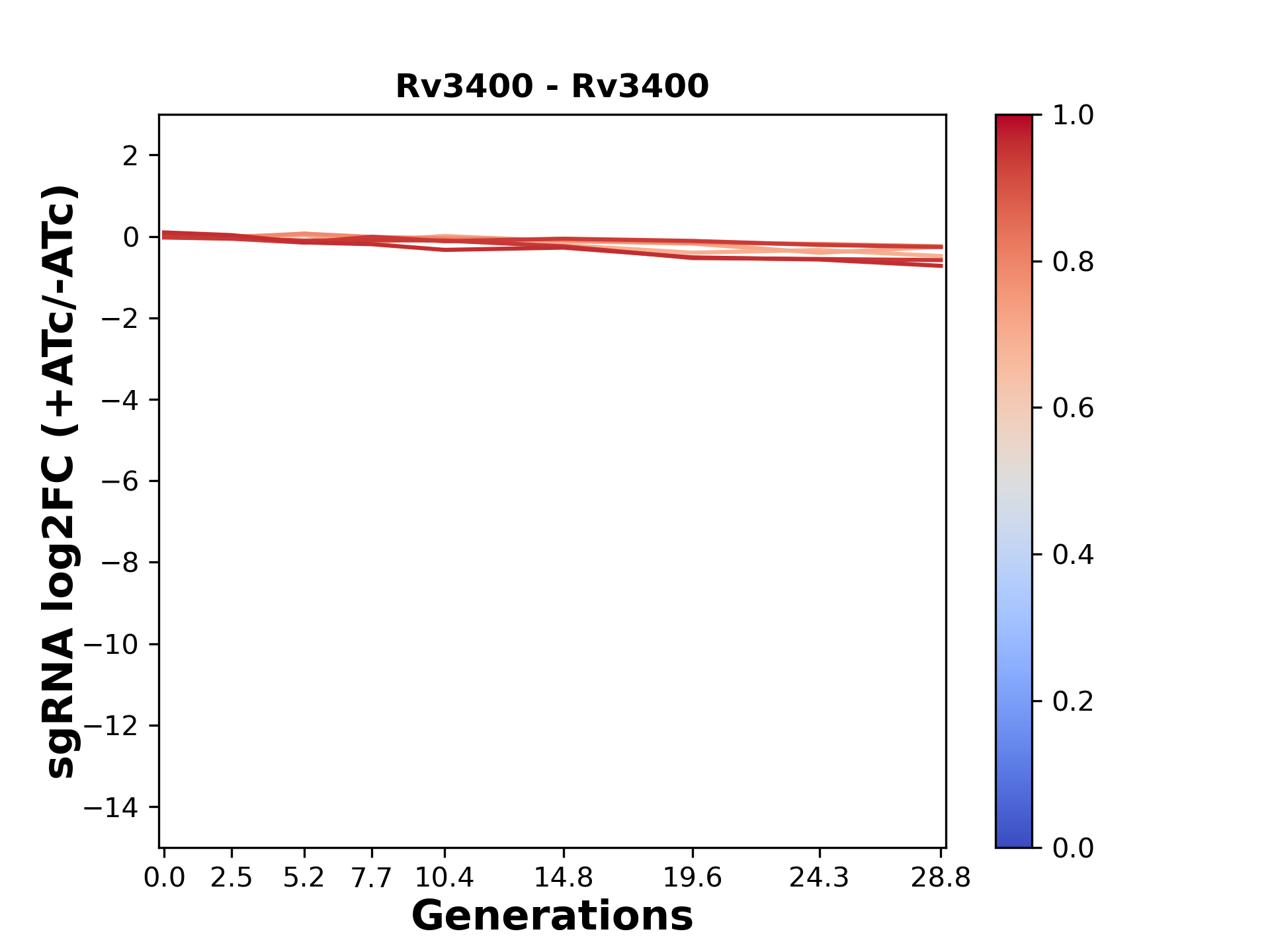
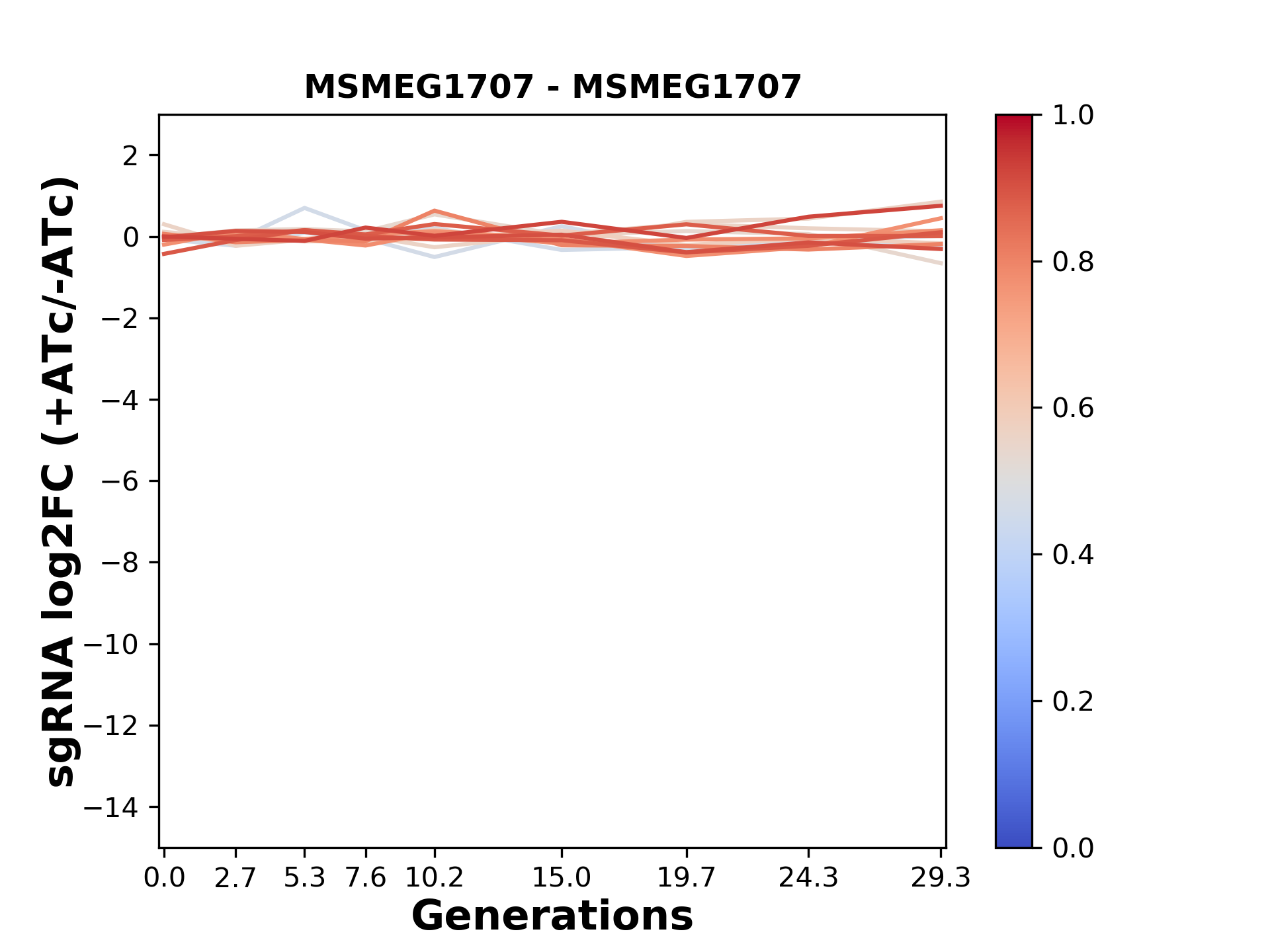
Here, we show sgRNA log2 fold-change values (L2FC) (plus versus minus ATc) over consecutive generations for sgRNAs targeting the gene you selected.
sgRNAs are color-coded based on their predicted strength from blue (strength=0; weak) to red (strength=1; strong).
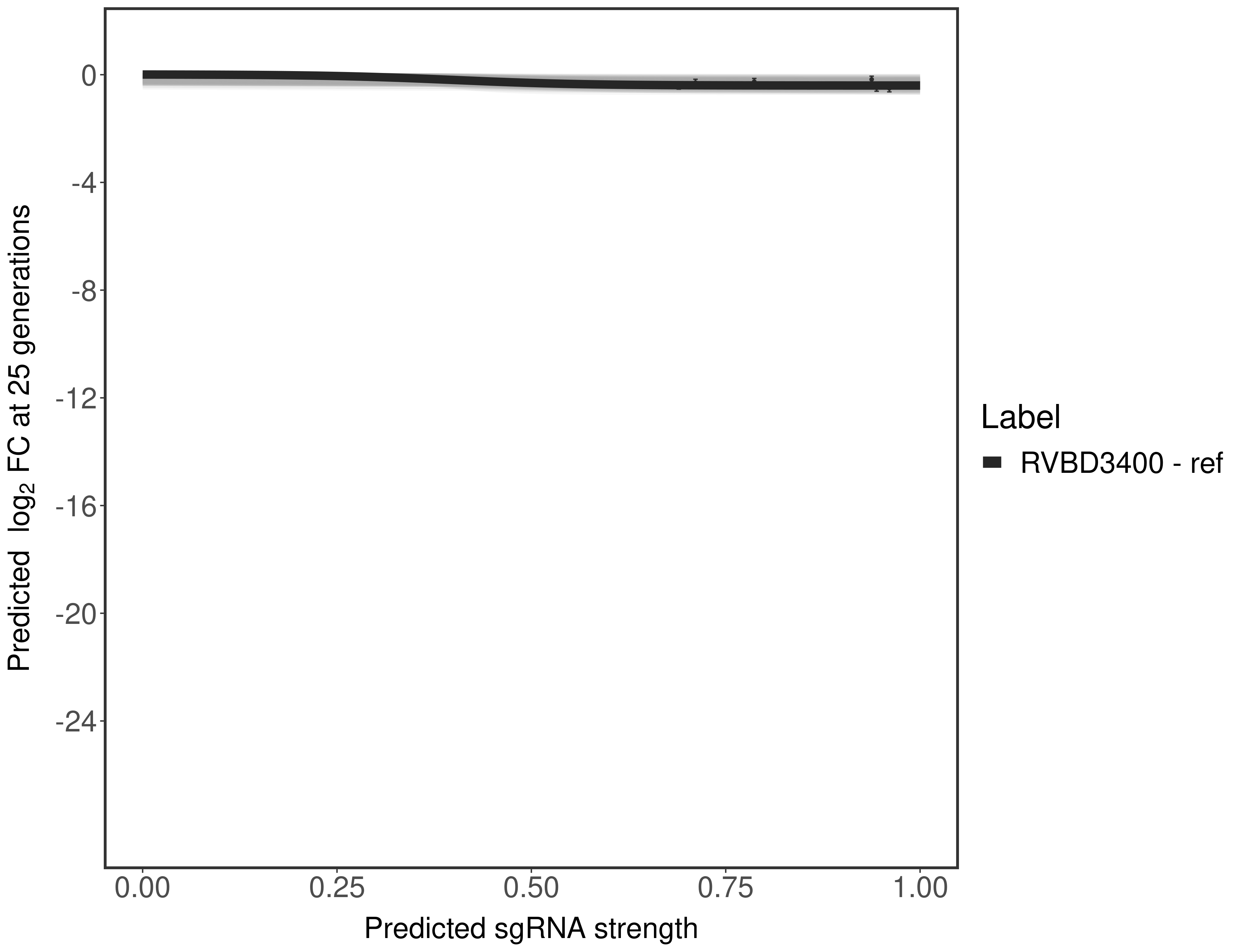
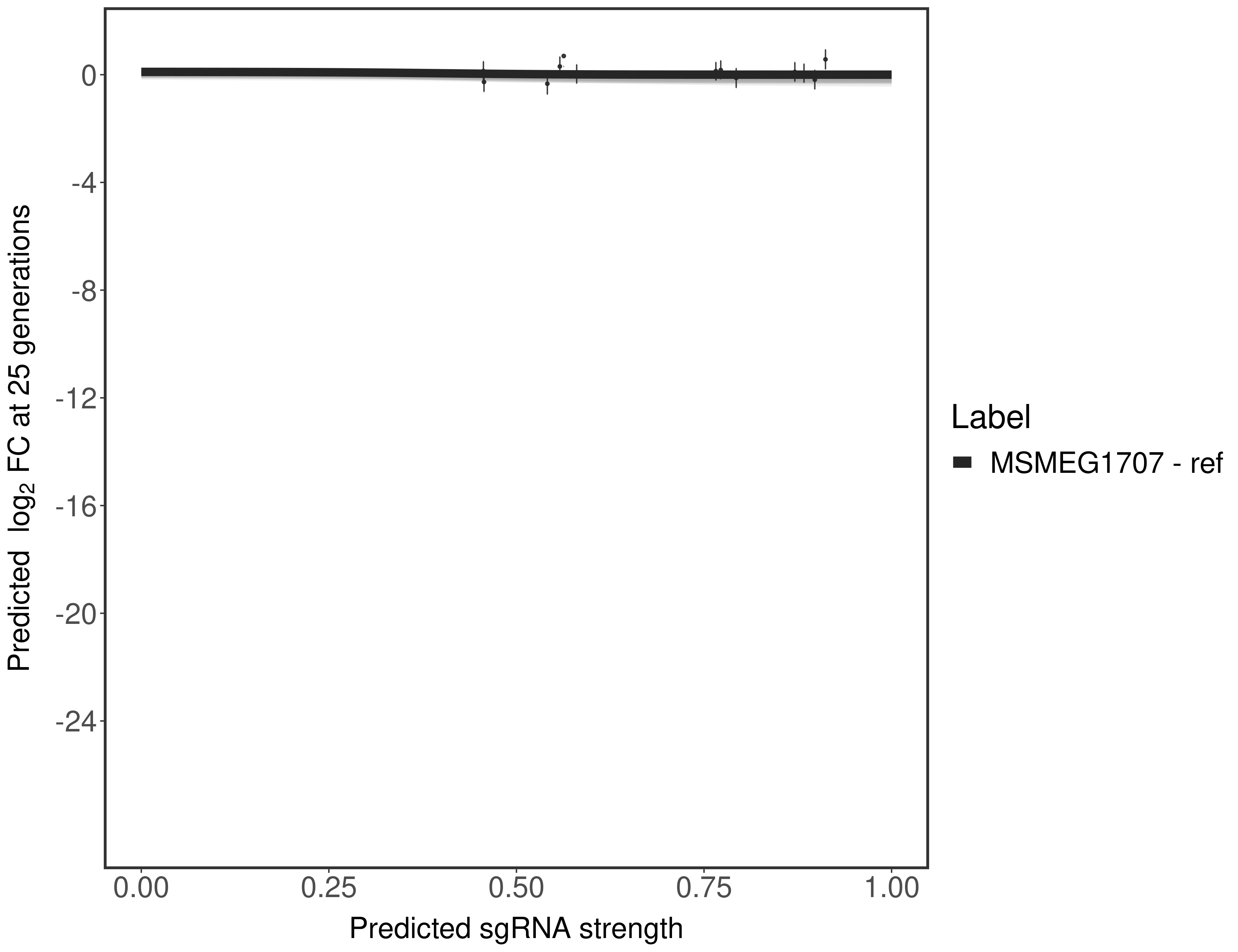
In order to quantify gene vulnerability, data were subsequently analyzed with a hierarchical model at the sgRNA level (two-line model) and gene level (Hill curve). The vulnerability quantification is shown below. [More information]
Experiment (PMID: 35637331): To define genes that control drug potency in Mtb, we performed 90 CRISPRi screens across nine drugs in the Mtb reference strain, H37Rv. These screens used a genome-scale CRISPRi library (RLC12) containing 96,700 sgRNAs to enable titratable knockdown for nearly all Mtb genes. Anhydrotetracycline (ATc) was added 1 or 5 days prior to drug exposure to transcriptionally activate CRISPRi and deplete target gene products. Drugs were screened at concentrations spanning the predicted minimum inhibitory concentration (MIC). Triplicate CRISPRi library cultures were outgrown for two weeks in the presence or absence of drug. After outgrowth, we harvested genomic DNA from cultures treated with three descending doses of partially inhibitory drug concentrations (High, Med, and Low) and analyzed sgRNA abundance by deep sequencing. Hits were identified by MAGeCK as those genes whose CRISPRi inhibition reduced or increased relative fitness in the presence of a given drug (false discovery rate (FDR) < 0.01, log2 fold-change |L2FC| > 1).
Shown below are the feature-expression heatmaps for this gene for the indicated drugs from the 1-day and 5-day CRISPRi library pre-depletion screens. The black triangles denote increasing drug dose (Low, Med, High). The color of each circle represents the gene-level L2FC. A white dot represents an FDR < 0.01 and a |L2FC| > 1. If the FDR for this gene under a given drug screening condition was greater than 0.01, no L2FC value is plotted.
You can click on each square in the feature-expression heatmap to display this gene's behavior in the presence of the indicated drug at the indicated dose in two volcano plots. The "sgRNA level" volcano plot shows the L2FC value and FDR for each sgRNA targeting the gene of interest. Only sgRNAs above the sequencing limit of detection (counts > 100 after normalizing for sequencing depth) are displayed. sgRNAs are color coded according to predicted strength (see Bosch & DeJesus et al. for details, PMID: 34297925). The "Gene level" volcano plot shows the L2FC value and FDR for each gene, with the gene of interest marked with a large green dot.
| 1 day predepletion | 
|
|---|---|
| 5 day predepletion | 
|
 |
You do not have permission to view notes.